
|
|
|
|
|---|---|---|---|
| RasMol script | Default RasMol display | ||
| Introduction |
Proteins, by their nature, have complex 3D structures which cannot always be easily appreciated from a 2D picture. A far better way of getting a feel and understanding of a protein's structure is to manipulate it interactively using a molecular graphics program.
The pricipal program we will be using throughout this course is RasMol. This is freely available and runs under Windows, unix, linux, and even on Macintosh/PPC computers. It allows you to rotate protein structures, zoom in on them, render them in different ways and using various colouring schemes, label atoms and residues, and so on.
Throughout this course you will see icons such as these:

|
|
|
|
|---|---|---|---|
| RasMol script | Default RasMol display | ||
These icons indicate that an example has been prepared for you for viewing in RasMol. Often these complement a 2D image illustrating a particular point and allow you to examine the 3D version in any way you like (ie from any angle, zooming in on any part, and so on) to get a better understanding of the point being made.
The first two of the above icons indicate a "RasMol script", whereas the third indicates RasMol's default depiction of the molecule. The distinction between these two is illustrated in the two examples below.
| Default RasMol display | RasMol script rendering |
|---|---|
|

|
| [Click on either image to enlarge] | |
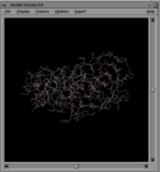
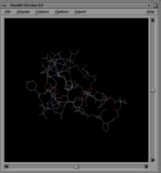
The first shows RasMol's default display mode for a protein structure. Here the covalent bonds between all atoms are displayed as thin lines, each half coloured according to the atom at that end (blue for nitrogen, red for oxygen, yellow for sulphur, white for carbon, etc). Apart from the bonds, little other detail about the structure is discernible.
In the second example the same protein is rendered to show its secondary structure in cartoon form. You immediately see there are two chains (one coloured purple and the other red), and both are all-beta-strand structures. What's more, there is a molecule bound between the two chains; it is shown in spacefill with its carbon atoms shown in black, nitrogens in blue and oxygens in red. Furthermore, the sidechains of the protein's crucial catalytic residues have been shown as thick sticks (just about visible here) interacting with the bound molecule.
The second image gives more information and gives it immediately. The rendering commands are defined using RasMol's scripting language and stored in the file that contains the coordinates. So when you click one of the RasMol script icons you will see the molecule immediately rendered so that the key point being made about its structure is immediately apparent.
For this to work, you not only need to obtain a local copy of the RasMol program, but you also need to configure your Web browser so that it knows when to invoke RasMol automatically and whether to call it in standard or script mode.
|
The aim of this page is to provide information on:
|
| Getting RasMol |
The following link explains how to download and install RasMol on your computer.

Getting & Installing RasMol 2.6

Getting & Installing RasMol 2.7- Recommended version as it displays multiple models
Once you have installed it on your computer try familarising yourself with the program. Below are two sets of instructions, the first a very basic introduction, and the second a more comprehensive operating manual.

Basic introduction to simple RasMol commands 
Online Operating Instructions
To help you get familiar with the program, the following link contains the 3D coordinates of a protein structure (in PDB format - which is just the format that RasMol reads). Save the file onto your local disk and load it into RasMol.
 PDB file (1a07.pdb)
PDB file (1a07.pdb)
The protein, in case you're interested, is the SH2 domain of a tyrosine kinase and consists of two chains and a bound ligand (see 1a07).
You may see or hear about a related program called Chime. This is a browser plug-in, rather than a stand-alone program, largely derived from RasMol. Some people prefer it over RasMol. However, Chime suffers a major shortcoming which makes it less useful for teaching: you cannot enter commands to select parts of your structure for separate rendering and examination. It does not have a "command window" as found in RasMol. Protein Explorer is a way of getting a command window for Chime using Java (a language for web browsers). Protein Explorer does do quite a lot more than this and makes easier access to one or two features of chime such as surfaces that are not in rasmol.
Chime suffers a number of further problems in that it is only properly supported on Windows and Macintosh and not on Unix. It works through your browser and so is dependent on your browser version and often new releases of browsers require revision of Chime to make them work. Protein Explorer is also dependent on the implementation of Java that comes with the browser, making it even more dependent than Chime on browser versions. The web pages for Protein Explorer advertise fixes for Unix and Linux, but they are not easy to do and involve buying extra software
What's more, once you have Chime installed, your browser will always favour it over RasMol as it is a plug-in, and, once installed, the program is fiendishly difficult to remove(!) - although see disabling Chime in Netscape 2 and 3.
So we strongly recommend you stick with RasMol for the duration of this course. We are able to give you advice if you have problems installing rasmol and interfacing it with the course. CHIME and Protein Explorer are too complicated and system dependent for us to help you.
Having said that, if you already have Chime installed on your computer and you can't (or don't want to) remove it, then don't worry. Chime is able to understand RasMol script files and will display them in exactly the same way. So you'll see exactly the same thing as you would in RasMol. The only point where you will get problems is where you need to use the command window to enter commands to operate on your structure (as in some of the practicals).
The best solution in these situations is, rather than click on the RasMol link, save the file behind it to disk instead. You do this by holding down the SHIFT key while you click on the link (or, under Windows, right-click on the link and choose the "Save to disk" option).
Once the file has been saved, run RasMol and, from the command window, load the file with either the
orload filename
script filename
command, depending on whether the file is a simple PDB or RasMol script file, respectively.
| Configuring your browser for RasMol |
You can configure your browser to automatically run RasMol and display a protein structure when you click on a "RasMol link".
The details on how you preform the configuration depend on which version of Netscape you are using, and whether yours is a Windows or unix/linux environment.
Also, you need to define two types of links: one for standard RasMol display, and the other for display of RasMol script files. Each type of link is defined by a "MIME" type. (MIME stands for Multimedia Internet Mail Enclosures. Basically it identifies the type of file being sent to your browser. You can configure your browser to let it know how to treat different MIME types).
In our case we want to define the following three MIME types:
MIME type Definition Action chemical/x-pdb Default RasMol display of PDB files Run RasMol chemical/x-ras Interpret and display RasMol script file Run RasMol with -script flag application/x-rasmol
The method of configuring your browser varies slightly from version to version. The following are instructions for the most common versions. If your machine or version of Netscape aren't represented here, choose the nearest and see if it enables you to perform the configuration.

Unix: Netscape versions 2 and 3 
Unix: Netscape version 4 
Windows: Netscape version 4 
Netscape version 7 and mozilla 
Internet Explorer on PCs (not recommended) 
Macintosh (not used at Birkbeck)
One problem you can have is if there is a space in the name of your temporary file directory this will cause rasmol to fail. C:\TEMP is fine c:\Temporary files\Nick Keep would not work. Om windows for netscape 4 and 7, the temporary file directory is controlled by the user environment variable TMP. To change this go to Control Panel, System, Advanced and select enviroment variables. Find TMP and edit it (you may want to change TEMP at the same time to the same address). Netscape 4 on Macs and Unix allow you to set this directory through preferences, applications. Netscape 7 on unix uses /tmp which should be on all machines anyway.
Some rasmol scripts contain the coordinates as well as the instuctions others just the instructions and have a 'load file.pdb' for the latter type of script to work you must already have looked at the PDB file by itself so that the coordinates are stored in your download directory which is where the script will look (ie the directory the script is in).
If none of the above seem to match your set-up, then try looking at the more detailed information available at:-

Chemical MIME
If you still get no joy, contact the course tutor.
| Testing your browser configuration |
Use each of the three links below to test whether your browser has been correctly configured. Click on each icon in turn. This should open a RasMol window in which you should see the image shown above the icon.
If all works OK, then you're all set to go. If not, check whether your configuration settings are correct. You can try downloading each file in turn and save to disk (ie by SHIFT-clicking on the icons below or, under Windows, right-clicking on them and then saving to disk when prompted). Then run RasMol and load the files as described above in "A note about Chime". If all fails, e-mail the course tutor.
| Default RasMol display | RasMol script rendering | ||
|---|---|---|---|
| chemical/x-pdb | chemical/x-ras | application/x-rasmol | |
|

|

|
[Click on any image to enlarge] |
|

|
|
|
|
Plant seed protein crambin from Abyssinian cabbage (PDB code 1crn) |
|||
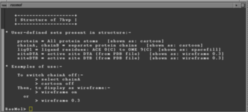
If your browser has been correctly configured, you will also get a RasMol "command window" displayed as follows:-
|
|---|
| [Click on image to enlarge] |
Note that, under Windows, the command window doesn't automatically pop up onto your screen when RasMol loads; instead, it is placed on the task bar at the bottom of the screen and you need to click on its task-bar icon to get it to spring into view.
You can use the command window to issue various commands for manipulating the objects displayed in the RasMol window (see documentation links above).
If there are any scripts in the course material that do not work after you have got the test scripts to work please email your tutor. There may be file extension variations used by other course authors we do not know about.